New Investigation Method Casts Light on Protein Synthesis
23 December 2011
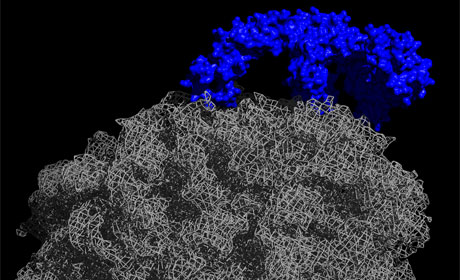
Figure: Center for Molecular Biology of Heidelberg University (ZMBH)
Heidelberg molecular biologists have achieved new insights into the synthesis of proteins with a newly developed innovative technique. A team of scientists headed by Prof. Dr. Bernd Bukau and Dr. Günter Kramer of the DKFZ-ZMBH Alliance in conjunction with Prof. Dr. Jonathan Weissman from the University of California in San Francisco devised this new method, which is called “selective ribosome profiling”. The Alliance is a research association between the Center for Molecular Biology of Heidelberg University (ZMBH) and the German Cancer Research Center (DKFZ). Selective ribosome profiling enables researchers to identify a cell’s protein synthesis profile. It was developed for a specific factor that assists in protein synthesis (molecular chaperone) to analyse the participation of chaperones in the folding of newly synthesised proteins. “These new insights provide the basis for a molecular understanding of how the factors involved in the maturation and folding of newly synthesised proteins are coordinated so that they do not get in each other’s way,” says Prof. Bukau. The findings have been published in the journal „Cell“.
Proteins are produced as long chains of amino acids that have to fold into a particular three-dimensional structure for the protein to function properly. Misfolding of proteins can damage cells and lead to neurodegenerative disorders like Parkinson’s disease. To guarantee correct protein folding, all cells possess a class of factors known as molecular chaperones. They assist in the folding of newly made proteins and ensure that the structure of existing proteins is maintained. However, the precise way in which chaperones assist in protein folding is still unclear.
With the aid of selective ribosome profiling, the scientists headed by Prof. Bukau and Dr. Kramer have now identified the protein synthesis profile of a cell and analysed the role played by the bacterial chaperone Trigger Factor in the folding of newly synthesised proteins. To this end, they isolated active ribosomes – the molecular machines responsible for making proteins – and used high-throughput sequencing to find out which protein was being produced at a particular time and whether Trigger Factor was supporting its folding. “For the first time, we were able to identify the entire range of newly synthesised proteins that are assisted by a chaperone in their folding,” says Dr. Kramer.
The scientists established that Trigger Factor makes contact with the majority of the cell’s proteins – well over 1,000 different species – already during synthesis. However, Trigger Factor does not bind every growing amino acid chain with the same efficiency. Instead, it chooses its binding partners largely on the basis of their future destination within the cell. “We were able to identify a new class of substrates bound more strongly by Trigger Factor than any of the substrates previously known,” says Dr. Kramer. The class in question is that of the pore proteins in the outer membrane of the bacterium Escherichia coli, which are largely responsible for the transport of substances into the cell.
Selective ribosome profiling also allows the precise determination of the point in time when the chaperone binds to the ribosome and the growing amino acid chain. “In contrast to assumptions held so far, Trigger Factor does not bind to the amino acid chain immediately as the latter emerges from the ribosome but only after a distinct time interval,” says Prof. Bukau. “This delay in binding results in a time frame for other factors to interact with the emerging amino acid chain, e.g. enzymes that also play an important part in protein maturation.” Although selective ribosome profiling was developed specifically for the chaperone Trigger Factor, it can be easily modified to investigate other factors from bacteria to humans. “Thus, we have laid an important foundation for the detailed investigation of the function of molecular chaperones in the folding of newly synthesised proteins,” says Prof. Bukau.
Original publication
E. Oh, A. Becker, A. Sandikci, D. Huber, R. Chaba, F. Gloge, R. Nichols, A. Typas, C. Gross, G. Kramer, J. Weissman, B. Bukau: Selective Ribosome Profiling Reveals the Cotranslational Chaperone Action of Trigger Factor in Vivo. Cell (2011), doi: 10.1016/j.cell.2011.10.044
Note for newsdesks
Digital picture material is available from the Press Office.
Contact
Prof. Dr. Bernd Bukau
Molecular Biology Centre (ZMBH) and German Cancer Research Centre
phone: +49 6221 546795
bukau@zmbh.uni-heidelberg.de
Communications and Marketing
Press Office
phone: +49 6221 542311
presse@rektorat.uni-heidelberg.de

