First Green, then Red – Fluorescent Dye Timer Tells the Age of Proteins
Joint press release of
Heidelberg University and
German Cancer Research Center in the Helmholtz Association
25 June 2012
In many diseases, from infections through to cancer, the protein metabolism in the cell is defective. Scientists from the German Cancer Research Center (DKFZ), the Center for Molecular Biology of Heidelberg University (ZMBH), and the European Molecular Biology Laboratory (EMBL) have now developed a method that enables them to monitor the aging process of proteins in a cell with unprecedented precision. The group has reported their results in the latest issue of Nature Biotechnology.
Proteins have important functions in our body: They confer structure, catalyze chemical reactions, serve as transport molecules for important substances, protect from pathogenic agents, and serve as an emergency source of energy. However, if the amount of a protein increases or decreases strongly, this often results in disease. If, for example, the p53 protein, which has been called the “guardian of the genome”, is broken down in an uncontrolled manner, processes like DNA repair, control of cell division, or induction of cell death cannot take place in the affected cell. As a result, the defective cell starts dividing uncontrollably and a tumor arises. To determine whether the protein metabolism of a cell is defective, researchers in the group of Professor Michael Knop have developed a novel method: They make proteins glow. However, instead of using a single fluorescent dye, as it is commonly done, the investigators have developed a complex of a red and a green fluorescent marker. This so-called tandem fluorescent protein timer (tFT) is linked to the protein during the very process of protein synthesis and, thus, delivers information about the amount, location and age of the molecules.
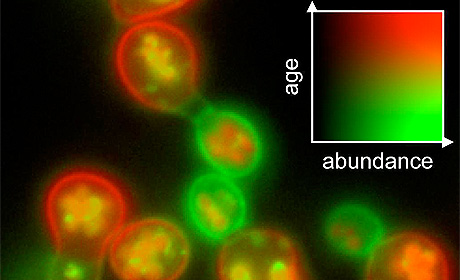
Michael Knop, leader of the Research Group “Cell Morphogenesis and Signal Transduction” in the DKFZ-ZMBH Alliance, explains how this new method works: “Immediately after the cell has formed the protein, the green dye – consisting of green fluorescent protein, GFP – starts emitting light. That means that in all those places in the cell where green light is emitted the molecule is found. Based on the color intensity of the emitted light, we are also able to determine the quantity of the protein.” From the fluorescence, the scientists are now also able to infer on the age of molecules. “As time progresses, the red fluorescent protein also starts emitting light. As a result, the proteins shift colors from green to red as they get older,” Knop continues to explain. “Thus, we can differentiate newly formed – green – proteins from old – red – ones. This enables us to calculate their lifespan and check whether a protein is being broken down more rapidly or more slowly than usual.” An additional advantage: tFTs produce very bright fluorescence so that the method has a high sensitivity.
The novel tFTs make it possible to monitor the age of proteins in a timeframe ranging from ten minutes up to several hours. If the green fluorescent protein (GFP) is combined with different fluorescent dyes, it is even possible to track breakdown processes taking place over several days. The investigators used yeast as a model organism. The reason: This single-celled organism is very similar to our cells in many basic processes and is therefore a suitable model. Moreover, Professor Elmar Schiebel, leader of the Research Group “Segregation of Chromosomes in Mitosis” in the DKFZ-ZMBH Alliance, has shown with his team that this method also works in human cells. This opens up whole new prospects for examining damaged cells and for developing new drugs to regulate protein stability in diseased cells.
Anton Khmelinskii, Philipp J Keller, Anna Bartosik, Matthias Meurer, Joseph D Barry, Balca R Mardin, Andreas Kaufmann, Susanne Trautmann, Malte Wachsmuth, Gislene Pereira, Wolfgang Huber, Elmar Schiebel & Michael Knop: Tandem fluorescent protein timers for in vivo analysis of protein dynamics. Nature Biotechnology, 2012, DOI: 10.1038/nbt.2281

