Heidelberg Researchers Study Formation of Bacterial Spores
8 March 2018
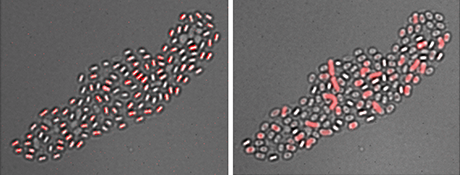
Picture: Alper Mutlu
Bacterial spores store information about the individual growth history of their progenitor cells, thus retaining a "memory" that links the different stages of the bacterial life cycle. This phenomenon was demonstrated in a recent study by an interdisciplinary research team led by Dr Ilka Bischofs at the BioQuant Centre of Heidelberg University. The spore memory could give rise to various adaptive behaviours in microbes. The results of the study were published in the journal "Nature Communications".
"To cope with fluctuations in nutrient availability, many bacteria can switch between two states," explains Dr Bischofs. "In the vegetative state, the cells grow and proliferate. Bacteria form spores to become dormant, which allows them to survive extended periods of starvation until new nutrients arrive to revive the spores." The researchers studied this adaptive bacterial life cycle using Bacillus subtilis as a model organism. Through the use of time-lapse microscopy, they were able for the first time in this context to observe and to study sporulation and spore revival at the single-cell level – and how they correlate. They discovered that the spores responded very differently to the influx of new nutrients: The spores that formed earlier during a nutrient down-shift revived more quickly.
The metabolic enzyme alanine dehydrogenase contributes to this effect, according to the researchers. Bacteria produce the enzyme when the amino acid L-alanine is available and stop synthesis once it runs out. Dr Bischofs explains that the enzyme is passed down from one generation of bacteria to the next by carry-over until spores are formed. The enzyme is then stored in the new spores, where it remains inactive until new nutrients arrive that facilitate spore revival and re-growth. "In this way, the spores obtain a stable phenotypic memory of the growth and gene expression history of their progenitor cells, which influences their future. The same basic principle could also apply to other cellular proteins."
The phenotypic memory has a major impact on spore development. The researchers were able to show a tradeoff between quantity and quality: Bacteria either make many spores that can revive only in nutrient-rich environments, or fewer but better spores that also revive in scarce environments. By linking the different phases of the bacterial life cycle, the spore "memory" could drive adaptations to ecological niches and trigger the emergence of various adaptive traits in microbes.
In addition to researchers at the Centre for Molecular Biology and the Institute for Pharmacy and Molecular Biotechnology of Heidelberg University, scientists from the German Cancer Research Center (DKFZ) and the Max Planck Institute for Terrestrial Microbiology in Marburg also joined in the study.

